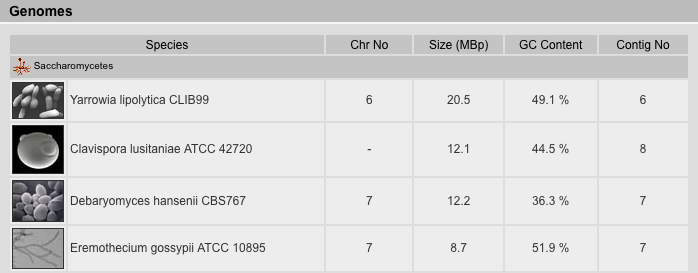
The Genomes Result View lists the species sorted by either taxonomy or name. It provides a table with several genome related data: (A) The number of chromosomes (if known). (B) The size of the genome given as number of base pairs included in the chromosome-, supercontigs-, or contigs-file, in descending priority. This is just a rough estimate because the chromosome- and supercontigs-files in most cases do not contain the unplaced supercontigs and contigs, respectively, which reduces the number of base pairs, but instead contain undefined bases "N" either in variable numbers (estimated e.g. from BAC-clone lengths) or by always including 50 or 100 bp which increases the number of bases. (C) the GC content in percent. (D) The number of entries in the highest priority file (chromosome > supercontigs > contigs).
